👥 Team








We are a research group in the Department of Computer Science at the University of Cincinnati, using AI to push the boundaries of healthcare. Our work spans immune checkpoint inhibitor discovery, spatial transcriptomics, cancer medical image analysis, and molecular simulation of biologics.
Our mission: make smart healthcare more personal, effective, and accessible — by building rigorous computational methods bridging machine learning and biomedical science.








MultiST jointly models spatial topology, gene expression, and H&E histomorphology through cross-attention fusion. Graph gene encoders with adversarial alignment + GAN–Fisher MMD latent refinement achieve sharper tumor microenvironment boundaries and more accurate therapeutic target localization. Wei Wang, Quoc-Toan Ly, Chong Yu, Jun Bai · arXiv 2601.13331 · 2026
Twin E(3)-invariant diffusion models generate pocket-aware peptide structures and amino-acid sequences de novo — leveraging geometric backbone representations and receptor pocket conditioning. Outperforms SOTA on binding success rate and structural similarity. Po-Yu Liang & Jun Bai · ISBRA 2025 (Springer LNBI)
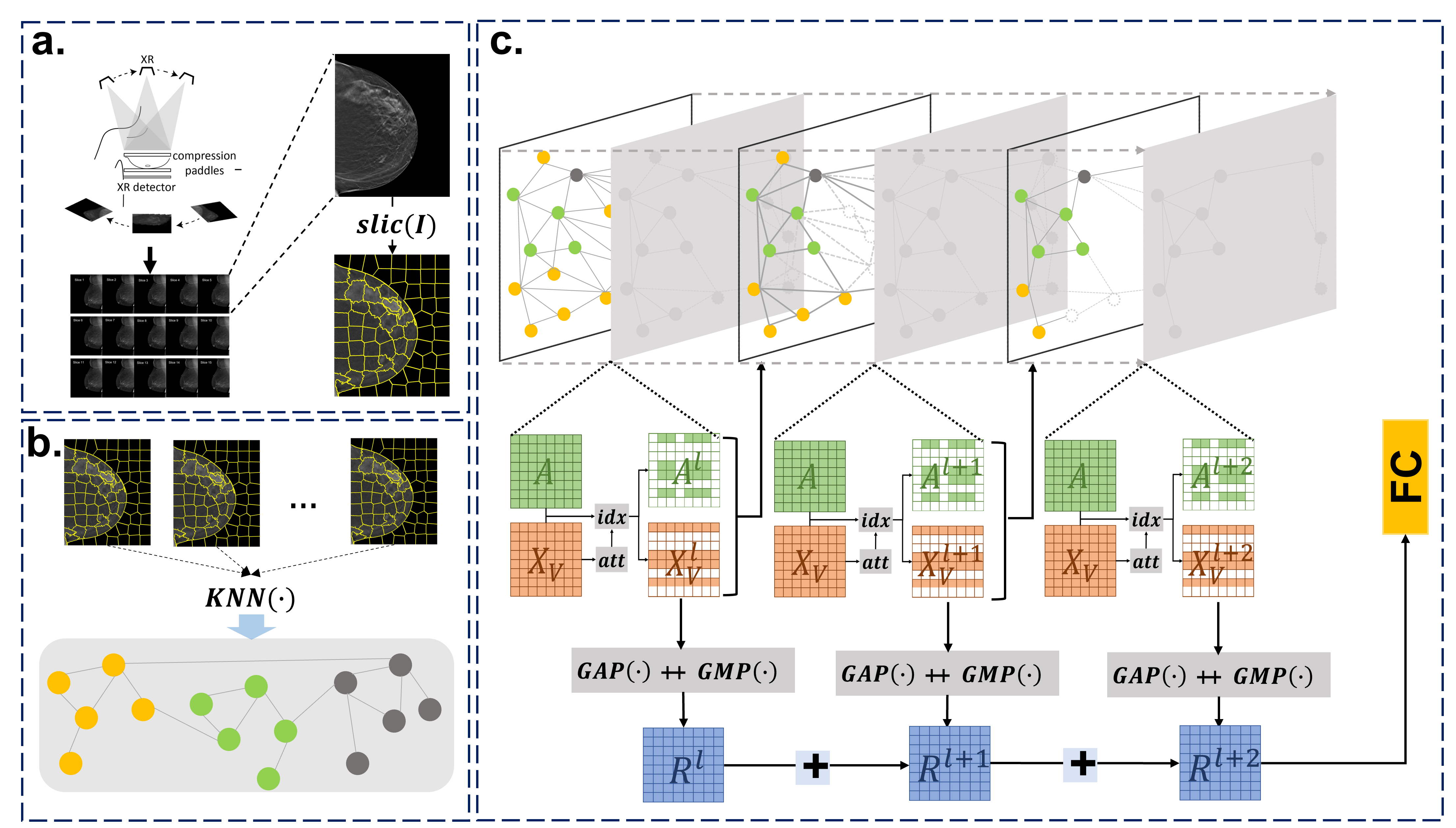
DBT volumes represented as graphs via SLIC + KNN, then classified by self-attention GCN with multi-scale GAP+GMP pooling — achieving state-of-the-art cancer detection and localization accuracy on 3D mammography screening datasets.
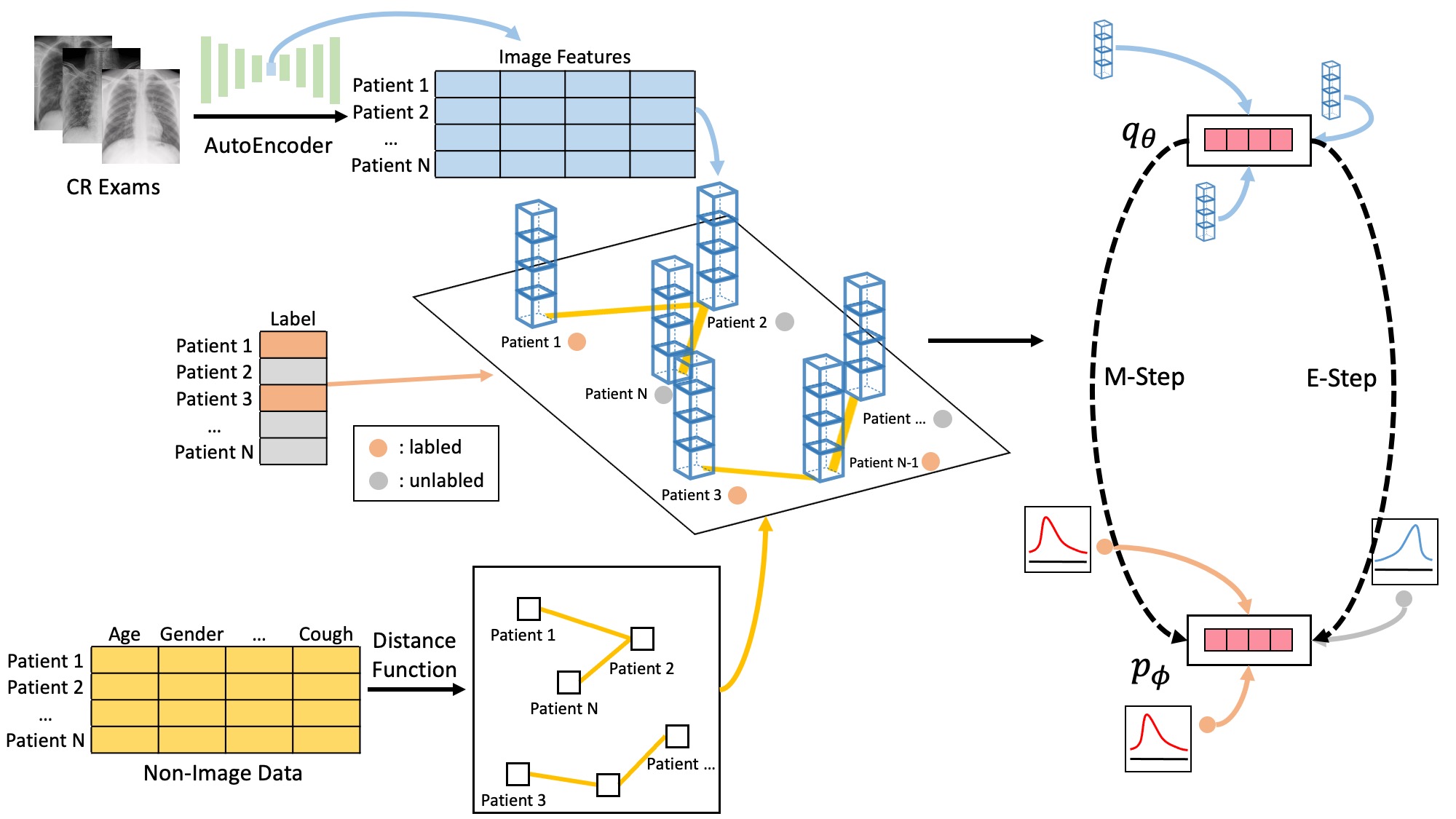
CGMNN fuses chest radiography autoencoder features with structured clinical data into a patient graph, classified by a Markov neural network using EM-based semi-supervised learning — providing actionable early ICU demand prognosis to optimize resource allocation.
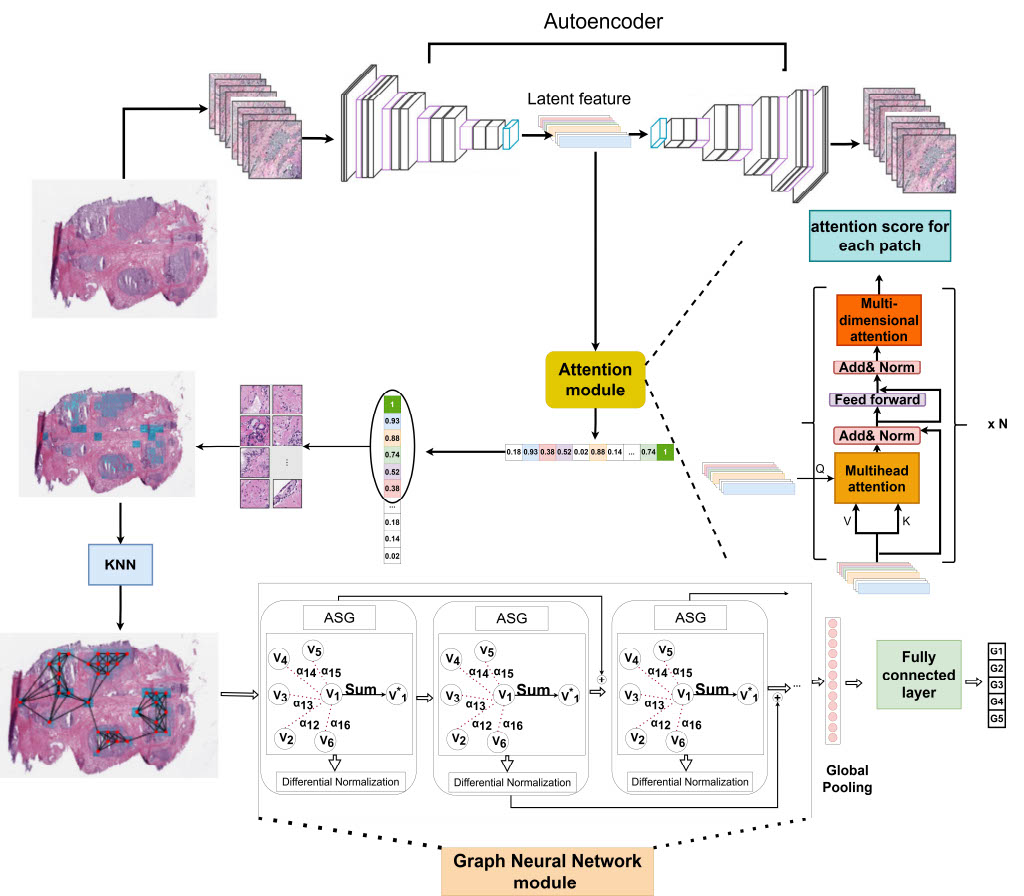
MIL-Transformer extracts discriminative patches from H&E WSIs, constructs a spatial patch graph, and applies gated-attention GCN for Gleason grade classification — no pixel-level annotation needed. Validated on TCGA-PRAD, PANDA & Gleason 2019 datasets.
Full list: Google Scholar ↗ | ResearchGate ↗
Core fundamentals of deep learning: architectures, optimization, CNNs, RNNs, transformers, and applications in science and engineering.
Introduction to programming concepts and computational thinking using Python.
Computing logic, algorithmic thinking, programming languages, and computing environments.
Data abstraction, functional abstraction, and problem solving in structured programming.
Data types, control structures, runtime environments, and programming paradigms.
Network types, topology, protocol architecture, routing, and performance.
Software lifecycle, development models, specification, design, and verification.
E/R modeling, relational theory, and practical DBMS experience.